The undidR package provides the framework for implementing difference-in-differences with unpoolable data (UNDID) developed in Karim, Webb, Austin, and Strumpf (2025). UNDID is designed to estimate the average treatment effect on the treated (ATT) in settings where data from different silos cannot be pooled together (potentially for reasons of confidentiality). The package supports both common and staggered adoption scenarios, as well as the optional inclusion of covariates. Additionally, undidR incorporates a randomization inference (RI) procedure, based on MacKinnon and Webb (2020), for calculating p-values for the UNDID ATT.
See the image below for an overview of the undidR framework: 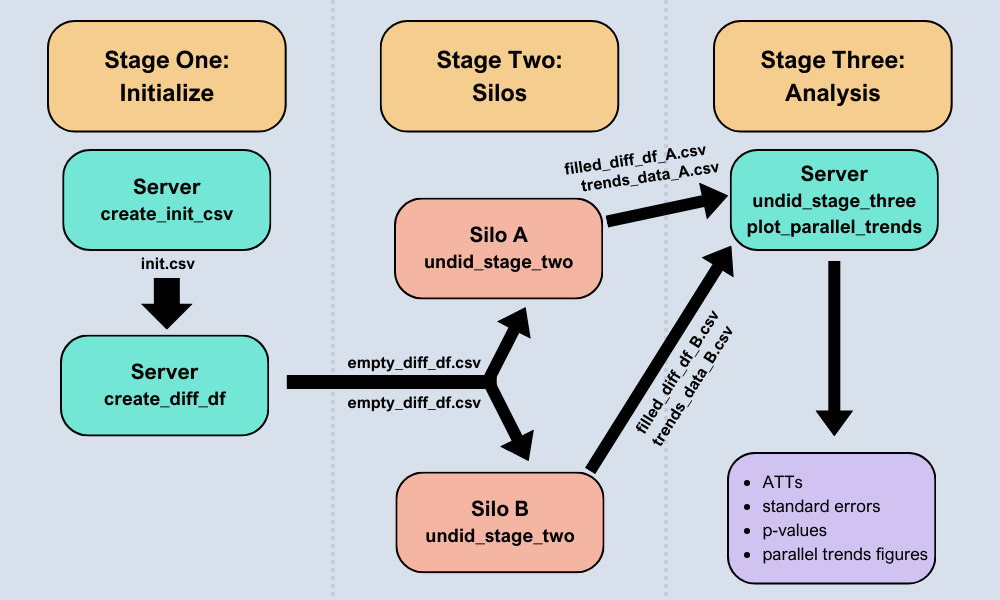
Installation
You can install the stable release version of undidR from CRAN with:
install.packages("undidR")Examples
For a set of detailed examples see the package vignette using:
vignette("undidR", package = "undidR")The following code chunks show some basic examples of using undidR at each of its three stages.
Stage One: create_init_csv() & create_diff_df()
library(undidR)
init <- create_init_csv(silo_names = c("73", "46", "54", "23", "86", "32",
"71", "58", "64", "59", "85", "57"),
start_times = "1989",
end_times = "2000",
treatment_times = c(rep("control", 6),
"1991", "1993", "1996", "1997",
"1997", "1998"))
#> init.csv saved to: C:/Users/Eric Bruce Jamieson/AppData/Local/Temp/RtmpAluOGo/init.csv
init
#> silo_name start_time end_time treatment_time
#> 1 73 1989 2000 control
#> 2 46 1989 2000 control
#> 3 54 1989 2000 control
#> 4 23 1989 2000 control
#> 5 86 1989 2000 control
#> 6 32 1989 2000 control
#> 7 71 1989 2000 1991
#> 8 58 1989 2000 1993
#> 9 64 1989 2000 1996
#> 10 59 1989 2000 1997
#> 11 85 1989 2000 1997
#> 12 57 1989 2000 1998
init_filepath <- normalizePath(file.path(tempdir(), "init.csv"),
winslash = "/", mustWork = FALSE)
empty_diff_df <- create_diff_df(init_filepath, date_format = "yyyy",
freq = "yearly",
covariates = c("asian", "black", "male"))
#> empty_diff_df.csv saved to: C:/Users/Eric Bruce Jamieson/AppData/Local/Temp/RtmpAluOGo/empty_diff_df.csv
head(empty_diff_df, 4)
#> silo_name gvar treat diff_times gt RI start_time end_time weights
#> 1 73 1991 0 1991;1990 1991;1991 0 1989 2000 both
#> 2 73 1991 0 1992;1990 1991;1992 0 1989 2000 both
#> 3 73 1991 0 1993;1990 1991;1993 0 1989 2000 both
#> 4 73 1991 0 1994;1990 1991;1994 0 1989 2000 both
#> diff_estimate diff_var diff_estimate_covariates diff_var_covariates
#> 1 NA NA NA NA
#> 2 NA NA NA NA
#> 3 NA NA NA NA
#> 4 NA NA NA NA
#> covariates date_format freq n n_t anonymize_size
#> 1 asian;black;male yyyy 1 year NA NA NA
#> 2 asian;black;male yyyy 1 year NA NA NA
#> 3 asian;black;male yyyy 1 year NA NA NA
#> 4 asian;black;male yyyy 1 year NA NA NAStage Two: undid_stage_two()
silo_data <- silo71
empty_diff_filepath <- system.file("extdata/staggered", "empty_diff_df.csv",
package = "undidR")
stage2 <- undid_stage_two(empty_diff_filepath, silo_name = "71",
silo_df = silo_data, time_column = "year",
outcome_column = "coll", silo_date_format = "yyyy")
#> filled_diff_df_71.csv saved to: C:/Users/Eric Bruce Jamieson/AppData/Local/Temp/RtmpAluOGo/filled_diff_df_71.csv
#> trends_data_71.csv saved to: C:/Users/Eric Bruce Jamieson/AppData/Local/Temp/RtmpAluOGo/trends_data_71.csv
head(stage2$diff_df, 4)
#> silo_name gvar treat diff_times gt RI start_time end_time weights
#> 1 71 1991 1 1991;1990 1991;1991 0 1989 2000 both
#> 2 71 1991 1 1992;1990 1991;1992 0 1989 2000 both
#> 3 71 1991 1 1993;1990 1991;1993 0 1989 2000 both
#> 4 71 1991 1 1994;1990 1991;1994 0 1989 2000 both
#> diff_estimate diff_var diff_estimate_covariates diff_var_covariates
#> 1 0.12916667 0.009655194 0.116348472 0.009930942
#> 2 0.06916667 0.008781435 0.069515594 0.008624581
#> 3 0.02546296 0.008134930 0.005133291 0.008084684
#> 4 0.02703901 0.008748277 0.029958108 0.008704568
#> covariates date_format freq n n_t anonymize_size
#> 1 asian;black;male yyyy 1 year 93 45 NA
#> 2 asian;black;male yyyy 1 year 98 50 NA
#> 3 asian;black;male yyyy 1 year 102 54 NA
#> 4 asian;black;male yyyy 1 year 95 47 NA
head(stage2$trends_data, 4)
#> silo_name treatment_time time mean_outcome mean_outcome_residualized
#> 1 71 1991 1989 0.3061224 0.1998800
#> 2 71 1991 1990 0.2708333 0.1502040
#> 3 71 1991 1991 0.4000000 0.1949109
#> 4 71 1991 1992 0.3400000 0.1876636
#> covariates date_format freq n
#> 1 asian;black;male yyyy 1 year 49
#> 2 asian;black;male yyyy 1 year 48
#> 3 asian;black;male yyyy 1 year 45
#> 4 asian;black;male yyyy 1 year 50Stage Three: undid_stage_three()
dir_path <- system.file("extdata/staggered", package = "undidR")
results <- undid_stage_three(dir_path, agg = "g", covariates = TRUE,
nperm = 399, hc = "hc0", verbose = NULL)
summary(results)
#>
#> Weighting: both
#> Aggregation: g
#> Not-yet-treated: FALSE
#> Covariates: asian, black, male
#> HCCME: hc0
#> Period Length: 1 year
#> First Period: 1989
#> Last Period: 2000
#> Permutations: 399
#>
#> Aggregate Results:
#> ATT Std. Error p-value RI p-value Jackknife SE Jackknife p-value
#> 0.0727396 0.02600099 0.04893262 0.05012531 0.03619008 0.06960867
#>
#> Subaggregate Results:
#> Treatment Time ATT SE p-value RI p-val JK SE JK p-val Weight
#> --------------------------------------------------------------------------------------------------------------
#> 1991 0.0434 0.0252 0.0899 0.3258 NA NA 0.2428
#> 1993 0.0478 0.0242 0.0528 0.4536 NA NA 0.2305
#> 1996 0.0451 0.0353 0.2106 0.6065 NA NA 0.0910
#> 1997 0.1322 0.0294 0.0001 0.0627 0.0532 0.0302 0.3863
#> 1998 -0.0812 0.0602 0.1937 0.2657 NA NA 0.0494
plot(results, lwd = 2, legend = "topleft")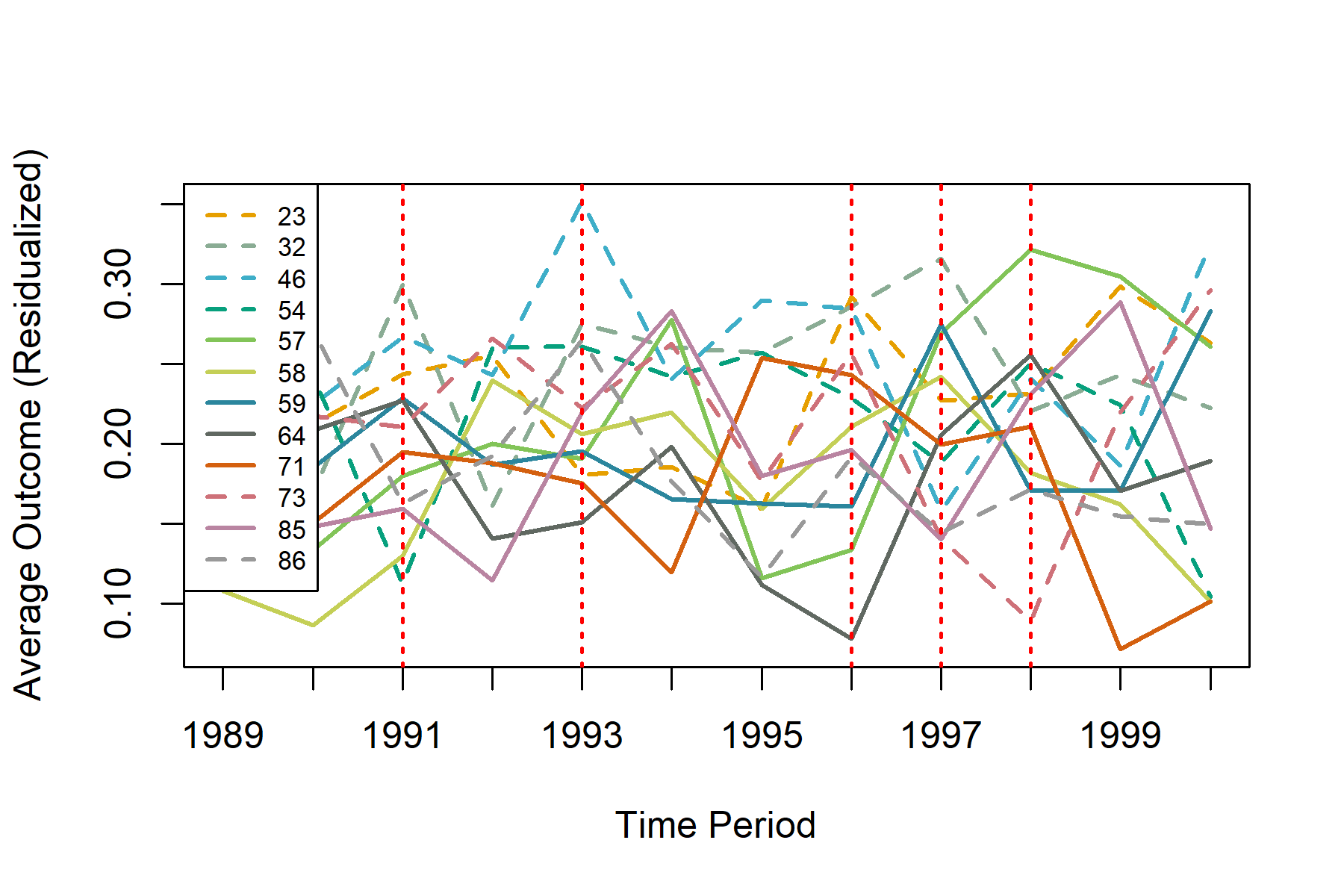
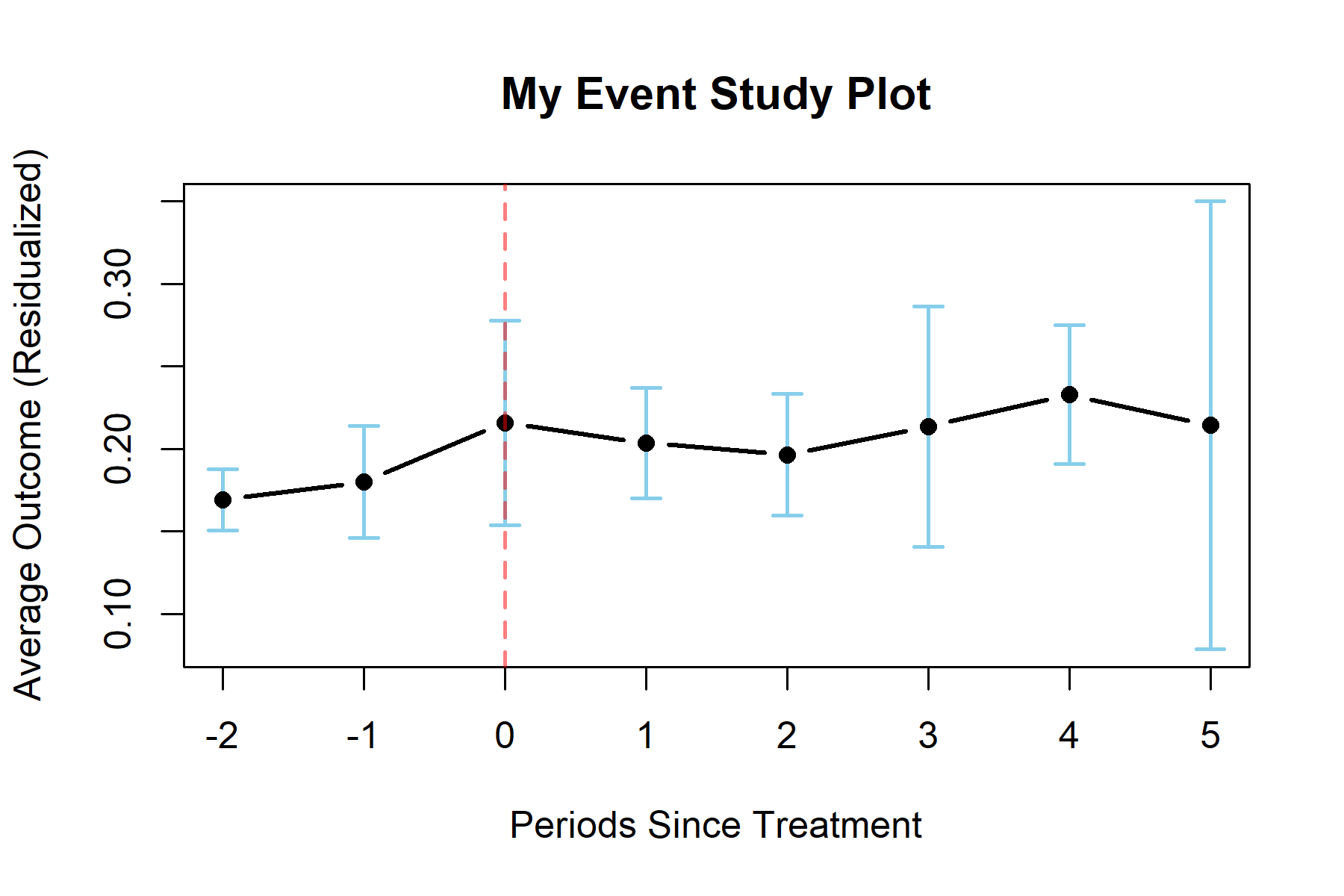
The S3 method of plot() for an UnDiDObj accepts parameters inherited from default plot() such as main, as well as the additional parameters of: ci (confidence interval), event_window (periods before / after treatment to plot), and event (TRUE or FALSE to plot either event study or parallel trends plots).
plot(results, event = TRUE, ci = 0.9, event_window = c(-2, 5),
main = "My Event Study Plot", lwd = 2)